Everyone is Launching AI Agents. What's Being Deployed?
A deep dive on the next cycle of biopharma's AI buildout
Last year's "growing buzz around AI agents" that we surveyed has since grown into a full avalanche of infrastructure commitments, partnerships, and agent launches across nearly every corner of biopharma. Let's take a fresh look.
A team at Stanford recently posted a preprint describing a system they called “Virtual Biotech”: a coordinated squad of AI agents organized to mirror a real drug discovery company, complete with a virtual Chief Scientific Officer, specialized scientist agents, and over 100 tools for querying biomedical databases.
For their headline demonstration, they deployed over 37,000 agents in parallel, each one tasked with annotating a single clinical trial, linking therapeutic targets to genomic and single-cell transcriptomic features. The resulting dataset spans 55,984 trials. The analysis turned up what the authors call previously unreported associations: drugs targeting cell-type-specific genes were 40% more likely to advance from Phase I to Phase II, 48% more likely to reach market, and showed 32% lower adverse event rates.
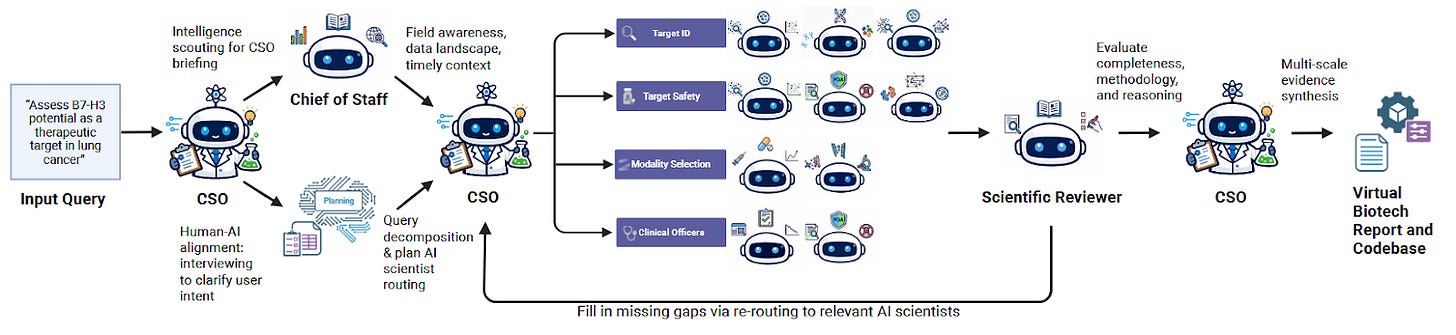
In another case study, the system pulled together genetics, transcriptomics, and clinical data on B7-H3 in lung cancer and landed on an antibody-drug conjugate strategy, the same bet several pharma companies are already running in the clinic. It also flagged liabilities and differentiation angles. The whole thing reportedly cost $46 in API credits and took less than a day.
The Virtual Biotech is one lab’s preprint, but it lands in what we recently likened to a “gold rush”—agents are being deployed across clinical operations, translational biology, antibody design, and regulatory workflows. Major pharma companies are in an apparent compute arms race, stacking GPU clusters and billion-dollar AI partnerships within months of each other. Startups backed by hundreds of millions are launching agent-focused platforms. NVIDIA’s Jensen Huang even went so far as to declare agentic AI “the new computer” at this year’s GTC.
Whether the implementations match is another question. In a recent experiment, researcher Liang Chang asked—”Can AI make better decisions than pharma executives?” and sent AI agent teams back to a pivotal 2012 decision in oncology, the BMS vs. Merck biomarker strategy that ultimately decided the Keytruda-Opdivo war, and found that both Claude and GPT independently recommended the same path BMS took. The path that lost.
The agents produced rigorous analysis, identified the exact competitive threat, and still followed the consensus. As Chang put it: “AI can give you the best possible analysis. It can’t give you the courage to go against it.”
What can AI agents do today, where are they falling short, and why is everyone building them?


