Cancer as a Data Problem: What AI Is Doing in Oncology
We track what has moved from promise to proximate execution, from AI-assisted candidate design with 2026 trial targets to agentic workflows that aim to handle multi-step oncology research tasks
Cancer can be looked at as a data problem because a tumor is an evolving population of cells, each accumulating mutations, signaling to neighbors, evading immune surveillance, adapting to treatment. The challenge of modeling has historically outrun the tools available to do it, but computers have been catching up.
Transformer architectures trained on biological data are beginning to predict drug response, generate therapeutic hypotheses, and identify which patients are likely to benefit from which treatments (part of a broader push that includes early attempts at virtual cell models) tasks that previously required years of wet-lab iteration. Some of that work is still early, though a handful of results have made it far enough through validation to be worth paying attention to.
Google Research, Google DeepMind, and Yale spent much of 2025 scaling C2S-Scale, a language model that reads single-cell RNA data as text; the 27-billion-parameter version, released in April, came in October with wet-lab validation of a model-generated hypothesis about making immune-”cold” tumors visible to T cells.
A collaboration between Microsoft Research, Providence Health, and the University of Washington took a complementary approach: GigaTIME, published in Cell in December, routinely converts pathology slides into virtual immune-protein maps, surfacing over 1,200 significant associations across 14,256 patients.
At Davos in January, Demis Hassabis now put Isomorphic Labs‘ first trials, primarily oncology candidates, at end of 2026; the company followed this month with IsoDDE, a general-purpose drug design engine that reportedly doubles AlphaFold 3’s accuracy, already deployed across its oncology programs.
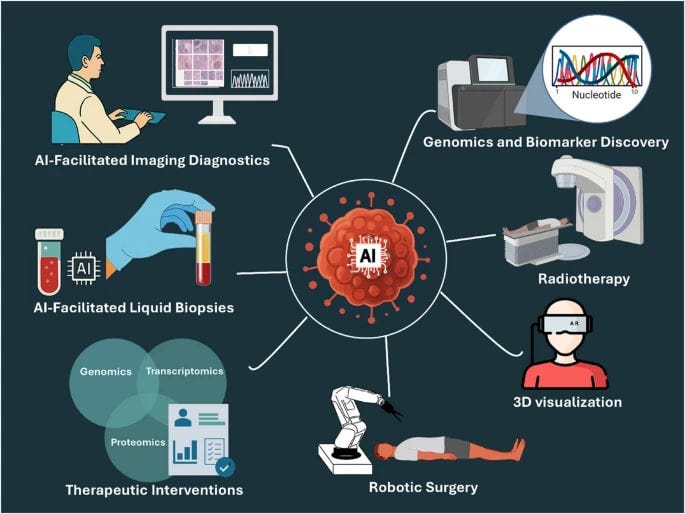
Not all of it is language-model work.